Characterization of eukaryotic DNA N6-methyladenine by a highly sensitive restriction enzyme-assisted sequencing | Nature Communications
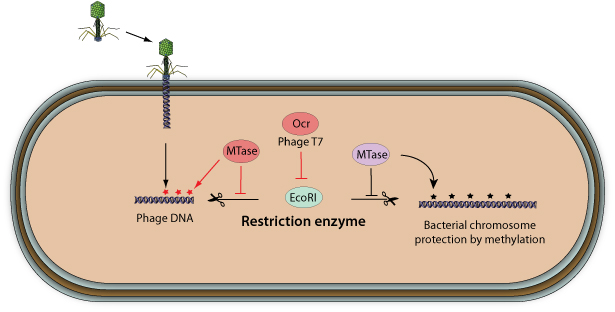
Why do restriction enzymes of bacteria not cut their own DNA? Why can it cut plasmids in genetic engineering, which are also part of the bacterial cell? - Quora
Restriction enzymes do not destroy the DNA of the same organism that makes them, because why? - Quora

Metagenomic methylation patterns resolve bacterial genomes of unusual size and structural complexity | The ISME Journal

REtools: A laboratory program for restriction enzyme work: enzyme selection and reaction condition assistance | BMC Bioinformatics | Full Text

Transient N4‐Acyl‐DNA Protection against Cleavage by Restriction Endonucleases - Jakubovska - 2019 - ChemBioChem - Wiley Online Library
A circular plasmid of 10, 000 base pairs (bp) is digested with two restriction enzymes A B, to produce a 3000 BP 2000 bp bands when visualised on an agarose gel. When
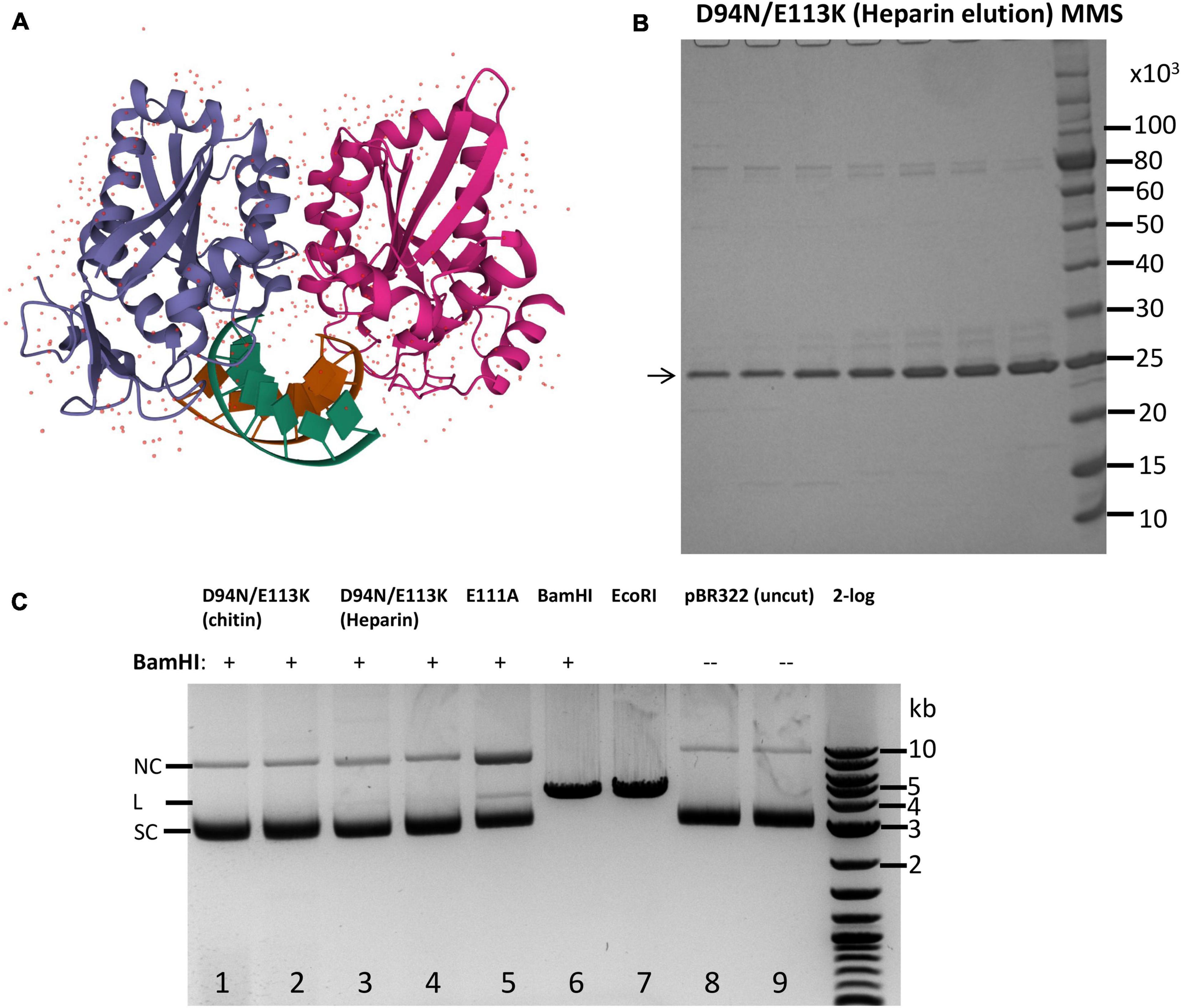
Frontiers | Engineering Infrequent DNA Nicking Endonuclease by Fusion of a BamHI Cleavage-Deficient Mutant and a DNA Nicking Domain